New paper out: functional metagenomics powered by synthetic biology
By EricvdHelm
Why do functional metagenomics and synthetic biology (synbio) make such an interesting combination? This week, our new article in Nature Chemical Biology ‘The evolving interface between synthetic biology and functional metagenomics’, sheds light on how progress in synthetic biology can advance, and already have advanced, the field of functional metagenomics.
We are facing a growing and aging world population, and mankind thus needs new drug molecules and ways to produce nutrients. Instead of using chemical synthesis, drugs and nutrients can be sustainably produced by modified bacteria. Moreover, most of those interesting molecules are already produced by billions of bacteria in the environment. Unfortunately, it is difficult to grow most types of bacteria in a laboratory, and it is therefore not possible to harness their useful capabilities directly. However, bacteria contain all the information needed to produce these valuable molecules in their DNA. Using methods known collectively as ‘functional metagenomics,’ the DNA of these bacteria can be recovered from the environment and used by host bacteria that can be cultivated in a lab. This allows us to make use of the capabilities of the billions of bacteria that are present in the environment without actually growing them, but by directly utilizing their DNA instead.
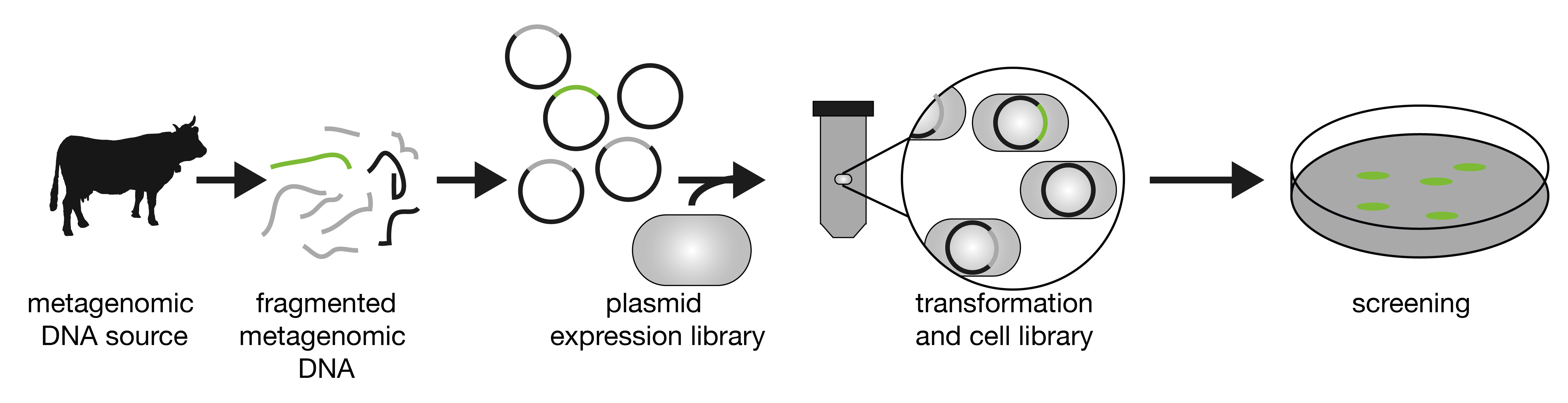 Construction of a metagenomic library. Environmental DNA is extracted, purified, fragmented and cloned into a shuttle vector. The library of plasmids is then transformed into an expression host such as Escherichia coli. Finally, the resulting clones can be analyzed according to their genotype and/or phenotype.
Construction of a metagenomic library. Environmental DNA is extracted, purified, fragmented and cloned into a shuttle vector. The library of plasmids is then transformed into an expression host such as Escherichia coli. Finally, the resulting clones can be analyzed according to their genotype and/or phenotype.
Which kind of metagenomics should be used?
In practice, there are two ‘metagenomic’ approaches, sequence-based approaches (where environmental DNA is sequenced and a function is assigned computationally) and function-based approaches (where the environmental DNA is transformed into a host bacteria and the genes are expressed and interrogated). In our article, we focused on the functional approach by specifically interrogating metagenomic DNA functionally using a genetic circuit.
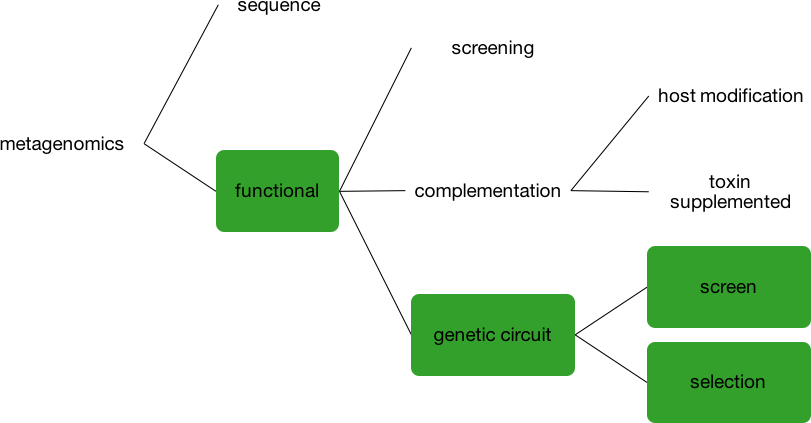
The term “metagenomics” can refer to many different techniques and procedures. In our new article, we focused on using genetic circuits to functionally mine a metagenomic library.
In our publication, we first surveyed the ways in which genetic circuits have been used in the recent past to interrogate metagenomic libraries. Though scientists have been quite creative, researchers need to move from a ‘screening’ method to a more high-throughput interrogation methods. These ‘screening’ methods require researchers to painstakingly examine each bacterial colony for a visual change associated with the production of a compound, for example. In more effective high-throughput methods, researchers couple the production of a compound of interest to the survival of the cell. The spontaneous death of cells lacking the target compound replaces the labor-intensive process of scrutinizing massive number of clones. My colleague Hans has utilized this approach previously to identify new vitamin transporters as outlined in the illustration below.
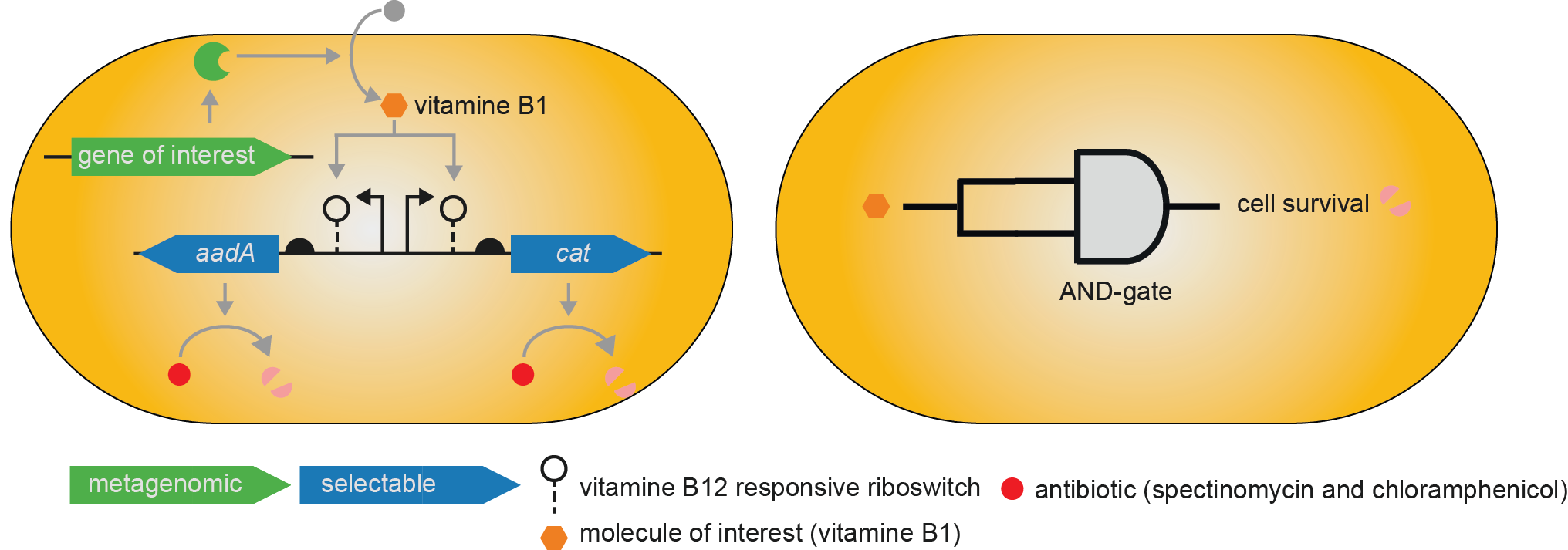
Example of a genetic circuit consisting of a riboswitch coupled with two selectable markers, which can be used to mine a metagenomic library for vitamin B1 transporting or producing genes. The genetic switch can also be formalized as an AND-gate with vitamin B1 as the input and cell survival as the output.
Insights for improving genetic selection circuits can also be obtained from biocontainment research, as it is notoriously difficult to perform experiments in which all cultured cells commit suicide. For example, the research group led by Farren Isaacs showed how multi-layered circuits can aid in this, and a recent review from the Collins lab summarizes the latest advances in biocontainment systems.
We anticipate that the expansion of synthetic biology tools, such as automated circuit design and computational design of proteins, will usher in greater efficiencies in the mining of functional metagenomics libraries. These advances in functional metagenomics and synthetic biology are already demonstrating remarkable potential in industrial and medical applications. Our full paper, available at Nature Chemical Biology, goes into more depth on all the previously constructed genetic circuits and new technologies that will continue to propel the field forward:
van der Helm, E. Genee, H.J. Sommer, M.O.A (2018) ‘The evolving interface between synthetic biology and functional metagenomics’ Nature Chemical Biology 10.1038/s41589-018-0100-x
Other resources
Note: parts of this blogpost are sourced from my PhD thesis
- functional metagenomics
- genetic circuit
- metagenomic library
- nature chemical biology
- synbio
- synthetic biology